สมการกรอบเคลื่อนที่ของเฟรชเนท์-เซอเรอท์ บนระนาบของระบบพิกัด ดี อาร์ พี ของการบิดตัวบนเส้นโค้งของโครงสร้างโปรตีน
Main Article Content
บทคัดย่อ
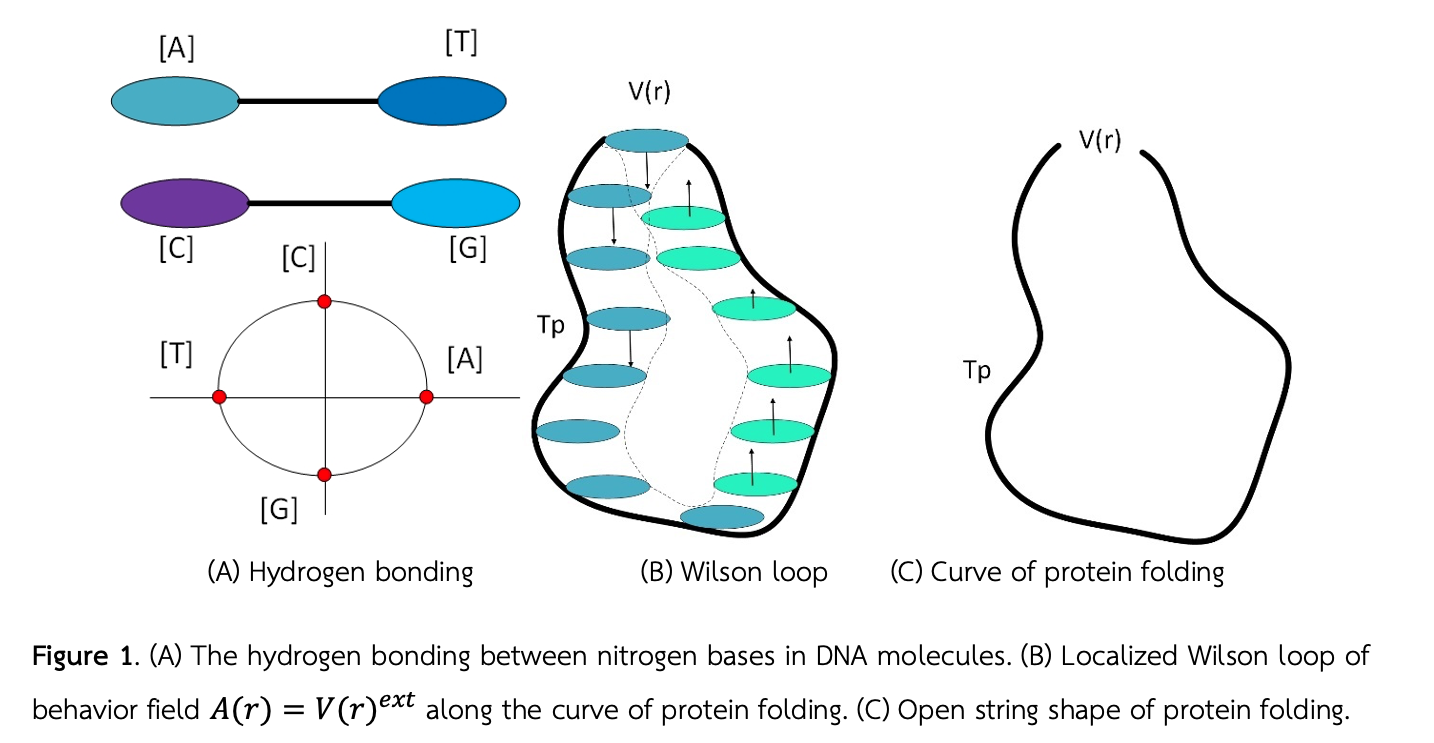
เราได้นิยามระบบพิกัดแบบกรอบเคลื่อนที่ได้บนเส้นโค้งหนึ่งมิติที่เป็นส่วนเกินของการบิดตัวการกาลเวลาและระยะทางบนระนาบการบิดตัวของโครงสร้างทุติยภูมิของโปรตีน งานวิจัยนี้คือส่วนการค้นพบทางทฤษฎีที่สำคัญที่สามารถตอบคำถามได้ว่าทำไมรูปร่างสามมิติของโครงสร้างของโปรตีนจึงมีลักษณะเหมือนกับเส้นเชื่อกปลายเปิดที่มีค่าความโค้งเป็นเส้นใยในระบบพิกัด (d,r,p)-layer เราได้เพิ่มคุณสมบัติพิเศษที่โครงสร้างทางคณิตศาสตร์ไปไม่ถึงเข้าไปเพิ่มเติมในสูตรของ in Frenet–Serret เพื่อใช้อธิบายคุณสมบัติการเข้าคู่กันและสนามพฤติกรรมแบบต่างๆของค่าจีโนไทป์ในสิ่งมีชีวิตที่เกิดจากระบบการเข้าคู่กันของโปรตีนที่และพฤติกรรมที่มีอยู่ในส่วนของ junk DNA และ non coding RNA ในโครโมโซม
Article Details
วารสารวิทยาศาสตร์และวิทยาศาสตร์ศึกษา (JSSE) เป็นผู้ถือลิสิทธิ์บทความทุกบทความที่เผยแพร่ใน JSSE นี้ ทั้งนี้ ผู้เขียนจะต้องส่งแบบโอนลิขสิทธิ์บทความฉบับที่มีรายมือชื่อของผู้เขียนหลักหรือผู้ที่ได้รับมอบอำนาจแทนผู้เขียนทุกนให้กับ JSSE ก่อนที่บทความจะมีการเผยแพร่ผ่านเว็บไซต์ของวารสาร
แบบโอนลิขสิทธิ์บทความ (Copyright Transfer Form)
ทางวารสาร JSSE ได้กำหนดให้มีการกรอกแบบโอนลิขสิทธิ์บทความให้ครบถ้วนและส่งมายังกองบรรณาธิการในข้อมูลเสริม (supplementary data) พร้อมกับนิพนธ์ต้นฉบับ (manuscript) ที่ส่งมาขอรับการตีพิมพ์ ทั้งนี้ ผู้เขียนหลัก (corresponding authors) หรือผู้รับมอบอำนาจ (ในฐานะตัวแทนของผู้เขียนทุกคน) สามารถดำเนินการโอนลิขสิทธิ์บทความแทนผู้เขียนทั้งหมดได้ ซึ่งสามารถอัพโหลดไฟล์บทความต้นฉบับ (Manuscript) และไฟล์แบบโอนลิขสิทธิ์บทความ (Copyright Transfer Form) ในเมนู “Upload Submission” ดังนี้
1. อัพโหลดไฟล์บทความต้นฉบับ (Manuscript) ในเมนูย่อย Article Component > Article Text
2. อัพโหลดไฟล์แบบโอนลิขสิทธิ์บทความ (Copyright Transfer Form) ในเมนูย่อย Article Component > Other
ดาวน์โหลด ไฟล์แบบโอนลิขสิทธิ์บทความ (Copyright Transfer Form)
เอกสารอ้างอิง
Andersen KG et al. (2020). The proximal origin of SARS-CoV-2. Nature Medicine, 1-3.
Borrie, M.S., Campor, J.S., Joshi, H. and Gartenberg, M.R. (2017). Binding, sliding, and function of cohesin during transcriptional activation. Proceedings of the National Academy of Sciences of the United States of America, 114(7), 1062-1071.
Byun, J.H., and Jung, I.H. (2018). Mathematical modeling of antibody drug conjugates with the target and tubulin dynamics to predict. Journal of Theoretical Biology, 443, 113-124.
Capozziello, S., Pincak, R. and Kanjamapornkul, K. (2017). Anomaly on superspace of time series data. Zeitschrift fuer Naturforschung A, 72(12), 1077-1091.
Capozziello, S., Pincak, R. and Kanjamapornkul, K. and Saridakis, E.N. (2018). Chern-Simons current in system of DNA-RNA. Annalen der Physik, 530(4), 1700271.
Cheng, J.X., Chen, L. et al. (2018). RNA cytosine methylation and methyltransferases mediate chromatin organization and 5-azacytidine response and resistance in leukaemia. Nature Communications, 9(1), 1163.
Cheng, W., Wang, S. et. al. (2018). C9ORF72 GGGGCC repeat-associated non-AUG translation is upregulated by stress through eIF2 phosphorylation. Nature Communications, 9, 02495.
Devi, M., Chingbiaknem, E. and Lyngdoh, R.H.D. (2018). A molecular mechanics study on GA codon box translation. Journal of Theoretical Biology, 441 28-43.
Einstein, A.,Podolsky, B. and Rosen, N. (1935). Can quantum-mechanical description of physical reality be considered complete? Physics Review, 47, 777-780.
Ghalami-Choobar, B. and Moghadam, H. (2018). Molecular docking based on virtual screening, molecular dynamics and atoms in molecules studies to identify the potential human epidermal receptor 2 intracellular domain inhibitors. Physical Chemistry Research, 6, 1154, 83-103.
Jaffe, R. L. (2005). Casimir effect and the quantum vacuum. Physics Review D, 72(5), 021301.
Jones, J. E. (1924). On the determination of molecular fields. -II. From the equation of state of a gas. Proceedings of the Royal Society A, 106(738), 463-477.
Kanjamapornkul, K. and Pincak, R. (2016). Kolmogorov space in time series data. Mathematical Methods in the Applied Sciences, 39(15), 4463-4483.
Kanjamapornkul, K., Pincak, R., Chunithpaisan, S. and Bartos, E. (2017). Support spinor machine. Digital Signal Processing, 70, 59-72.
Zhang, L. (2013). The van Der Waals force and gravitational force in matter. arXiv:1303.3579.
Merkenschlager, M. and Nora, E.P (2016). CTCF and Cohesin in Genome Folding and Transcriptional Gene Regulation. Annual Review of Genomics and Human Genetics, 17, 17-43.
Merkenschlager, M. and Odom, D.T. (2013). CTCF and cohesin: Linking gene regulatory elements with their targets. Cell, 152(6), 1285-1297.
Muangchoo-in, K., Sitthithakerngkiet, K., Sa-Ngiamsunthorn, P. and Kumam, P. (2021), Approximation theorems of a solution of amperometric enzymatic reactions based on Green’s fixed point normal-S iteration. Advances in Difference Equations, 2021(1), 1-13.
Nora, E.P. et al. (2020). Molecular basis of CTCF binding polarity in genome folding. Nature Communications, 11(1), 5612.
Ohtsuki, T. (2017). On the asymptotic expansions of the Kashaev invariant of hyperbolic knots with seven crossings. International Journal of Mathematics, 28(13), 1750096.
Peck, J.R. and Waxman, D. (2018). What is adaptation and how should it be measured? Journal of Theoretical Biology, 447 190-198.
Peters JM., Tedeschi A. and Schmitz, J. (2008). The cohesin complex and its roles in chromosome biology. Genes & Development, 22(114), 2389.
Pincak, R.,Kanjamapornkul, K. and Bartos, E. (2019). Forecasting Laurent Polynomial in the Chern-Simons Current of V3 Loop Time Series. Annalen der Physik, 531(7),1800375.
Pincak, R., Kanjamapornkul, K. and Bartos, E. (2020). Cohomology theory for biological time series. Mathematical Methods in the Applied Sciences, 43(2), 552-579.
Pincak, R.,Kanjamapornkul, K. and Bartos, E. (2020). A theoretical investigation on the predictability of genetic patterns. Chemical Physics, 535, 110764.
Rezazadegan, R. and Barrett, C. and Reidys, C. (2018). Multiplicity of phenotypes and RNA evolution. Journal of Theoretical Biology, 447 139-146.
Richterova, J., Huraiova, B. and Gregan, J.(2017). Genome Organization: Cohesin on the Move. Molecular Cell, 66(4), 444-445.
Saad, N.Y. (2018). A ribonucleopeptide world at the origin of life. Journal of Systematics and Evolution, 56(1), 1-13.
Saldana-Meyer, R. et al. (2019). RNA Interactions Are Essential for CTCF-Mediated Genome Organization. Molecular Cell, 76(3), 412-422.e5.
Shen, Z., Asa, S.L. and Ezzat, S. (2018). The retrotransposon gag domain containing protein Rgag4 is an Ikaros target in the pituitary. Molecular and Cellular Endocrinology, 461, 188-193.
Shinde, P., Sarkar, C. and Jalan, S. (2018). Codon based co-occurrence network motifs in human mitochondria. Scientific Reports, 8(1), 3036.
Soslau, G. (2018). Circular RNA (circRNA) was an important bridge in the switch from the RNA world to the DNA world. Journal of Theoretical Biology, 447, 32-40.
Vedani, A. (1988). YETI: An interactive molecular mechanics program for small molecule protein complexes. Journal of Computational Chemistry, 9(3), 269-280.
Witten, E. (2011). Fivebranes and Knots. arXiv:1101.3216v2.
Witten, E. (2012). Khovanov Homology and Gauge Theory. Argxrv:1108.3103v2.
Witten, E. (2016). The parity anomaly on an unorientable manifold. Physical Review B, 94(19), 195150.
Weyl, H. (1921). Feld und Materie. Annalen der Physik, 65(14), 541-563.
Wood, K.E. and Komarova, N.L.S. (2018). Cooperation-based branching as a mechanism of evolutionary. Journal of Theoretical, 445, 166-186.